 APPENDICES
APPENDICES APPENDICES
APPENDICESAppendix I.1. The lac operon control circuit.
To feed E. coli growth, glucose undergoes a series of biochemical transformations known as glycolysis (literally, “sugar breaking”) followed by oxidation through the Krebs cycle. These transformations produce both precursors for building other cell molecules and high-energy compounds that provide the chemical fuel for a wide variety of essential cell activities. To metabolize lactose for growth, E. coli utilizes both glycolysis and the Krebs cycle but requires additional biochemical capabilities to convert lactose into glucose. These activities include an enzyme (LacZ) that splits lactose into glucose and galactose and a cell membrane protein (LacY) that transports lactose into the cell (Jacob and Monod 1961). While the enzymes for glucose utilization are present under most growth conditions, LacZ and LacY appear only when the bacteria grow on lactose (Monod and Cohn 1952). Thus, their synthesis is regulated according to nutritional conditions, and expression of the corresponding DNA sequences lacZ and lacY is subject to a control circuit that has been largely deciphered over the past five decades.
When lactose utilization was first studied genetically, in the late 1940s and early 1950s, mutations affecting it clustered as a single point on the genetic map of E. coli:
![]()
As biochemical and genetic
analysis advanced, Monod et al. found that lacZ
and lacY constitute a contiguous structure in
the E. coli DNA, together with lacA
that encodes the coregulated acetylase
protein,
which detoxifies aromatic beta-galactosides from plants. They
called the entire genetic structure “the lac
operon” because it operates as a coordinated system (Jacob, Perrin et
al. 1960). By 1961, the operon was known to
contained four coding sequences (lacI encoding
repressor, lacZ encoding beta-galactosidase,
lacY encoding galactoside permease, and lacA encoding galactoside acetylase) plus a
regulatory site, lacO or the “operator,”
which controlled expression of lacZYA (Jacob and Monod
1961):
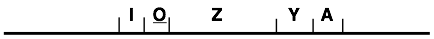
Over the next two decades, additional control sites and regulatory proteins were identified: the CRP protein (cyclic AMP receptor protein) and its cognate CRP binding site, the lacP promoter which was the starting site for transcription, and two additional operator sites, one of which resides inside the lacZ coding sequence (Scaife and Beckwith 1966; Arditti, Scaife et al. 1968; Silverstone, Magasanik et al. 1969; Muller, Oehler et al. 1996).
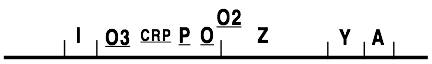
Expressing the lacZYA operon requires two proteins: (i) RNA polymerase that transcribes the DNA into a messenger RNA for further translation into LacZ and LacY proteins by the cell’s ribosomes and (ii) CRP (“cyclic AMP receptor protein”) which stimulates the initiation of transcription by the polymerase (Inada, Kimata et al. 1996; Kimata, Takahashi et al. 1997; Kuhlman, Zhang et al. 2007). Whether or not expression occurs is under the control of three additional molecules: (iii) the LacI repressor (I for “inducibility”) Willson, 1964 #964, (iv) an allolactose inducer molecule produced from lactose by the action of LacZ (Huber, Wallenfels et al. 1975; Huber, Kurz et al. 1976; Friedman, Olson et al. 1977; Turner and Huber 1977; Wu, Bahl et al. 1978) and (v) cyclic AMP (cAMP), a derivative of ATP that must bind to CRP before it can stimulate transcription (Inada, Kimata et al. 1996; Kimata, Takahashi et al. 1997).
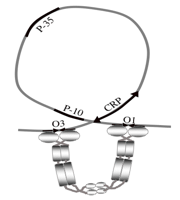 A key point is that the system works properly because lac operon DNA contains specific short binding
sequences for LacI repressor (“operators” = lacO),
for RNA polymerase (“promoter” = lacP) and
for CRP (“crp site”). When there is no lactose
(and hence no inducer), LacI binds to a pair of lacO
sites and prevents RNA polymerase from gaining access to lacP, where it has to bind to initiate
transcription of lacZ and lacY.
When inducer is present, LacI binding to lacO
sites no longer blocks polymerase access. By itself,
polymerase recognizes lacP weakly, and
physiologically significant transcription from lacP
requires simultaneous binding of a complex between cAMP and
CRP to the crp site (De Crombrugghe,
Chen et al. 1971; De Crombrugghe, Chen et al. 1971; Nisseley, Anderson et al. 1971; Pastan and Perlman 1972). Interaction
between CRP and polymerase stabilizes binding to lacP
and thus enhances transcription (Savery, Rhodius et
al. 1996). In the absence of cAMP, CRP
cannot bind to crp. These binding sites for
protein recognition (lacO, lacP and crp) must be properly positioned for all these
regulatory interactions to occur correctly. In essence, they
format the DNA for regulation and controlled access to genomic
protein coding information.
A key point is that the system works properly because lac operon DNA contains specific short binding
sequences for LacI repressor (“operators” = lacO),
for RNA polymerase (“promoter” = lacP) and
for CRP (“crp site”). When there is no lactose
(and hence no inducer), LacI binds to a pair of lacO
sites and prevents RNA polymerase from gaining access to lacP, where it has to bind to initiate
transcription of lacZ and lacY.
When inducer is present, LacI binding to lacO
sites no longer blocks polymerase access. By itself,
polymerase recognizes lacP weakly, and
physiologically significant transcription from lacP
requires simultaneous binding of a complex between cAMP and
CRP to the crp site (De Crombrugghe,
Chen et al. 1971; De Crombrugghe, Chen et al. 1971; Nisseley, Anderson et al. 1971; Pastan and Perlman 1972). Interaction
between CRP and polymerase stabilizes binding to lacP
and thus enhances transcription (Savery, Rhodius et
al. 1996). In the absence of cAMP, CRP
cannot bind to crp. These binding sites for
protein recognition (lacO, lacP and crp) must be properly positioned for all these
regulatory interactions to occur correctly. In essence, they
format the DNA for regulation and controlled access to genomic
protein coding information.
The level of cAMP in the E. coli cell is controlled by the availability of glucose in the external medium (Inada, Kimata et al. 1996; Kimata, Takahashi et al. 1997). Since cAMP is necessary for significant levels of lac operon transcription, it connects the regulatory circuit to recognition of one of the two sugars. The glucose control of cAMP levels occurs through one of the glucose transport proteins at the cell membrane (Harwood, Gazdar et al. 1976; Peterkofsky 1988). When glucose is available, the transport protein transfers a negatively charged phosphate group to the entering sugar, thus trapping it inside the cell and feeding it into the glycolytic process. This transport molecule is named “Protein IIAGlc” of the phosphotransferase system (PTS) discovered by Saul Roseman in 1964 (Kundig, Ghosh et al. 1964; Roseman 1969). The superscript “Glc” indicates that this particular PTS protein II specifically recognizes glucose (Simoni, Levinthal et al. 1967; Stock, Waygood et al. 1982). When there is no glucose available outside the cell, this protein accumulates in a form carrying an attached phosphate group, and this phosphorylated transport protein stimulates the enzyme that converts ATP into cAMP. Via this series of interactions, the cAMP levels inside the cell inversely reflect the glucose levels outside the cell. The glucose transport protein thus serves a dual function in both trapping glucose inside the cell and sensing extracellular glucose. Note, in a similar vein, that through the series of reactions leading up to derepression, LacY, LacZ and LacI repressor constitute a sensory circuit for lactose.
All of the reactions just summarized can be restated in a small number of Boolean (if X, then Y; or X ==> Y) propositions to compute the following specific and nontrivial regulatory algorithm: transcribe lacZ and lacY from lacP if, AND ONLY IF, glucose is NOT available and lactose IS available:

Since this formulation of lac operon control circuits was elaborated
several years ago, there has been a challenge to the role of
cAMP and CRP in establishing diauxic growth (Crasnier-Mednansky
2008; Narang 2009; Narang 2009). The
alternative explanation for glucose utilization prior to
lactose consumption is the observation that glucose is able to
inhibit transport of lactose into the cell by low uninduced
levels of LacY permease. That interpretation fails to explain
the observation that addition of cAMP abolishes the diauxic
growth pattern (Perlman and Pastan
1968; De Crombrugghe,
Perlman et al. 1969; Perlman, De Crombrugghe et al. 1969; Pastan and Perlman 1970). It may well
be that the two regulatory effects reinforce each other, in a
“fuzzy logic” manner (Kosko and Isaka 1993).
Appendix
I.2.
S. cerevisiae pheromone
response signal network (Sprague
1991). 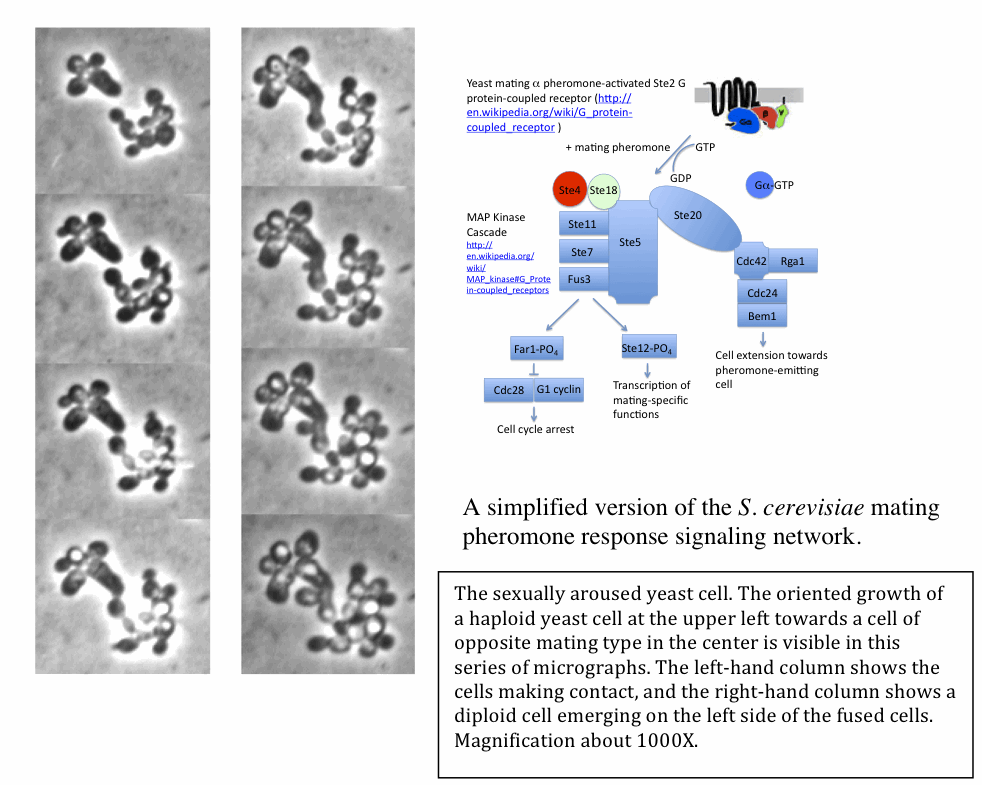
Appendix II.1. Formatting the genome for
transcriptional control.
Transcriptional control signals bind a wide variety but finite number of different regulatory proteins, or “transcription factors” (Hickey, Conway de Macario et al. 2002; Wang and Dohlman 2006; Hughes and Friedman 2007; Pinney, Amoutzias et al. 2007; Haerty, Artieri et al. 2008; Shan, Zahn et al. 2009; Gramzow, Ritz et al. 2010). Some of these factors are specialized, like the LacI repressor protein; others are more general and regulate families of interconnected functions, like the CRP protein; and other proteins are used by a significant proportion of all coding sequences, like RNA polymerase and (in eukaryotes) associated “basal transcription factors.”
The distribution of the corresponding recognition signals through the genome falls correspondingly into a range of classes: some unique to one or a small number of loci, like lacO operators; some present at several to many sites, like the crp binding sites; and some very widespread, like the promoter sequences that direct RNA polymerase to the initiation site for transcription (Jaeger, Chan et al. 2010). Even widespread signals, like promoters, can fall into more specialized classes; these classes facilitate the expression of coordinated suites of coding sequences required under special sets of conditions, such as high temperature “heat shock” and other stresses (Wallrath, Lu et al. 1994; Arsene, Tomoyasu et al. 2000; Pirkkala, Nykanen et al. 2001; Hengge-Aronis 2002; Klauck, Typas et al. 2007). Special promoter signals are also utilized for the expression of complex functions requiring transcription of many coding sequences, such as cell differentiation. In prokaryotes, which have highly condensed genomes, transcription signals are often found within the protein coding sequences. For example, one of the three lacO operators is located within the lacZ coding sequence.
Transcriptional control signals operate in combinations that facilitate regulatory interactions between the corresponding transcription factors. Several key regions of coding sequences hold the binding sites: the promoter area around the start site for transcription, the area around the termination site for ending transcription, and more distant locations that interact with the promoter area through DNA looping to bring them into proximity (Tjian 1995). These distant regions have received a variety of names (‘enhancer.” “silencer,” “upstream activating sequence”) according to whether they stimulate or repress transcription, but the term “enhancer” has found the most general application (Grimm and Nordeen 1999; Veitia 2008). Enhancers contain clusters of binding sites and are generally functional wherever they are located within distances of thousands or sometimes tens of thousands of base-pairs from the regulated coding sequences. In eukaryotes, which have interrupted coding sequences, enhancer elements, are often found in introns, which are spliced out of the final protein-coding messenger RNA (mRNA).
Transcription control signals in eukaryotes assemble hierarchically into larger complexes that have been given the generic description of “cis-regulatory modules” (CRMs) (Ben-Tabou de-Leon and Davidson 2007; Hinman, Nguyen et al. 2007; Bonn and Furlong 2008; Ben-Tabou de-Leon and Davidson 2009; Ho, Johnsen et al. 2009; Li and Davidson 2009; MacArthur, Li et al. 2009; Zinzen, Girardot et al. 2009; Borok, Tran et al. 2010). These CRMs can extend for many thousands of basepairs in the genome and typically contain multiple enhancer regions that are independently active at different times in the cell cycle, at different phases or tissues of multicellular development, or in response to distinct stimuli. This hierarchical architecture facilitates the execution of complex regulatory algorithms in morphogenesis and of multivalent responses to inputs, such as DNA damage or binding of intercellular signaling molecules (Cooper, Loose et al. 2009). By analogy with the multivalent allosteric nature of proteins, the composite CRMs constitute a type of microprocessor element. Although protein microprocessors can function at different locations in the cell and thoughout the genome, the action of CRMs is typically limited to nearby regions of the genome and the coding sequences they contain. In combination, CRMs and the transcription factors they bind can generate an enormous variety of genomic responses to input signals.
Transcription factors are dynamic proteins that regulate the expression of other transcriptions (and of themselves, as well) through binding to the CRMs adjacent to their coding sequences. These feedback interactions and the networks they form permit cells to construct complex regulatory circuits that process a large number of inter- and intracellular signals to compute an appropriate transcriptional response (Mangan and Alon 2003; Amoutzias, Robertson et al. 2004; Levine and Davidson 2005; Davidson and Erwin 2006; Madan Babu, Teichmann et al. 2006; Alon 2007; Balaji and Aravind 2007; Madan Babu, Balaji et al. 2007; Gonzalez Perez, Gonzalez Gonzalez et al. 2008; Aravind, Anantharaman et al. 2009; Hinman, Yankura et al. 2009). How these transcription factor circuits operate has been partially deciphered in certain model systems, such as the developing sea urchin, the progressive segmentation of the early Drosophila embryo and the establishment of anterior-posterior (AP) and dorsal-ventral (DV) axes in animal development (Nusslein-Volhard and Wieschaus 1980; Davidson and Erwin 2006; Erwin and Davidson 2009) and (De Robertis, Oliver et al. 1990; Nusslein-Volhard 1996).
Appendix II.2. Natural genetic engineering in B lymphocytes for the rapid evolution and maturation of a virtually infinite diversity of antigen-binding proteins.
Antibody structure. To clarify the discussion, it is necessary to describe some essential features of the antibody proteins and introduce basic terminology. I apologize to the reader for the proliferation of symbols, but that is inevitable when we have to deal with a system that has a virtually infinite number of outcomes. With a little patience, the description will all make sense and show how infinity is achieved in the real world of the B lymphocyte.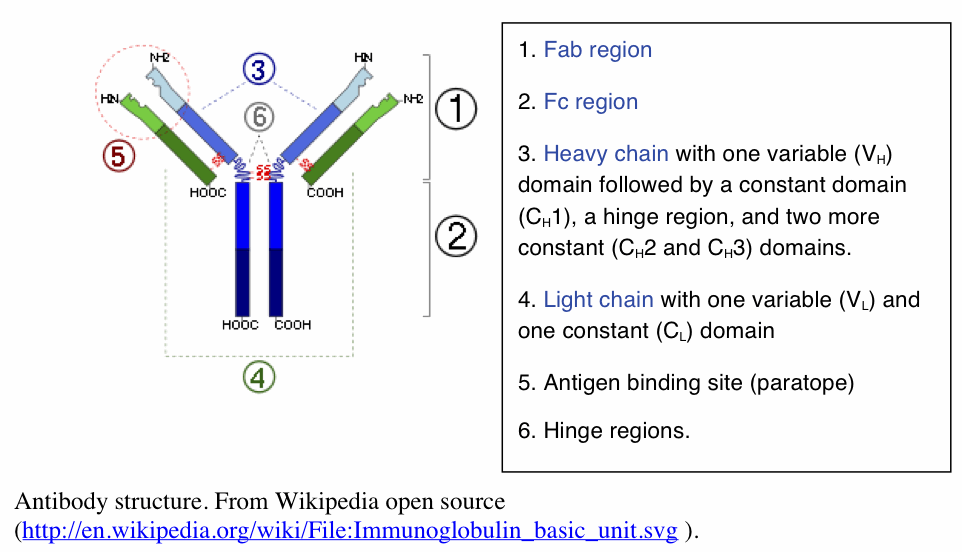
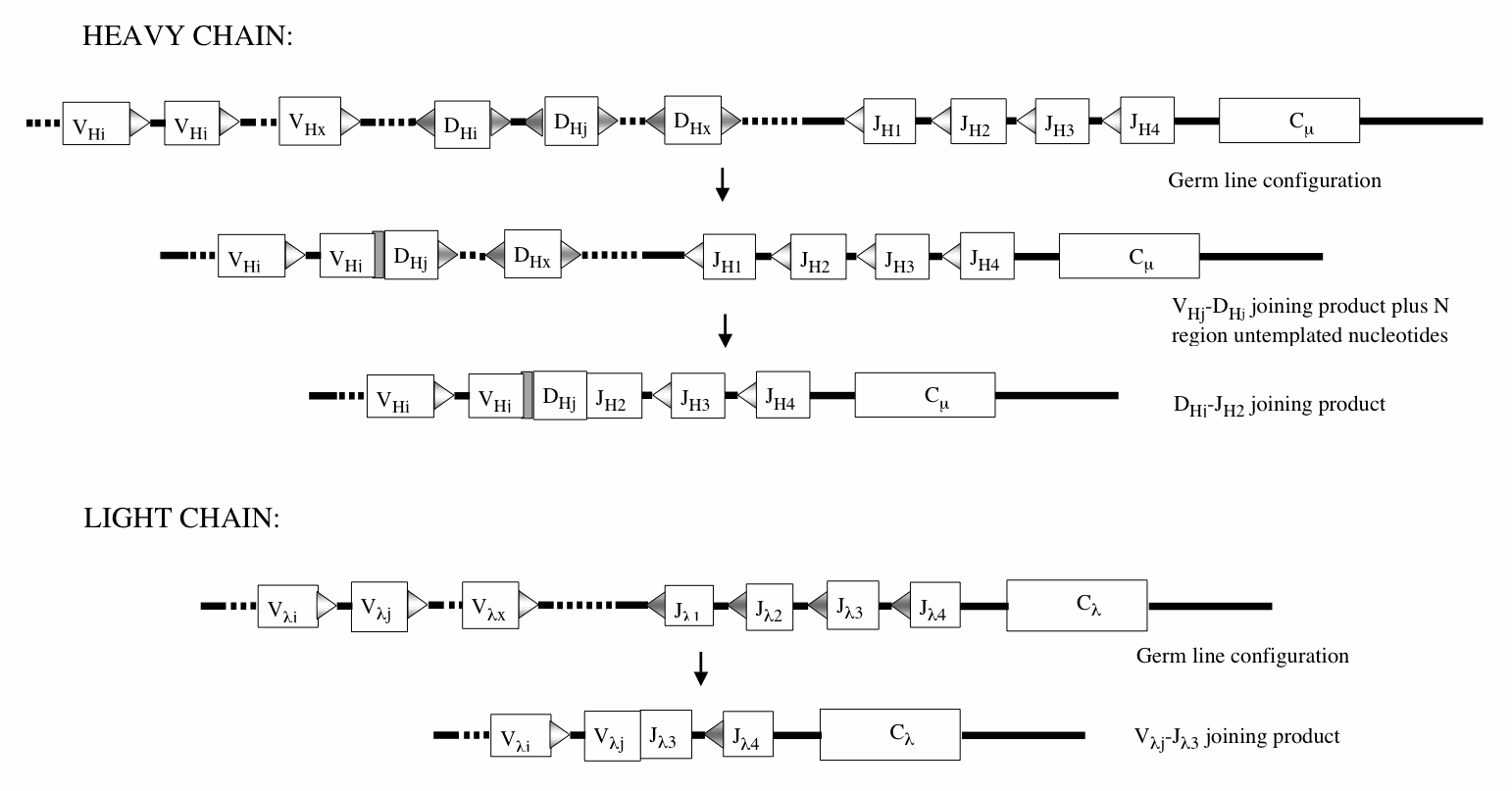
Structure of immunoglobulin-coding DNA and the process of V(D)J joining. The different V, D, J and C exons and the details of the molecular events are explained in (Bassing, Swat et al. 2002)and (Gellert 2002). The differently shaded triangles represent complementary recombination signal sequences (RSSs). For any two exons to join together, they must be flanked by complementary RSSs. Two identical RSSs will not promote DNA breakage and rejoining. Note, in the heavy chain chromosome, how the arrangement of RSSs prevents V-J and D-D joining and effectively prevents further joining activity on the fully rearranged chromosome. The shaded rectangle in the VHj-DHj joining product indicates a segment of “N region” untemplated nucleotides arising from the action of terminal transferase before the broken fragments are ligated together. From (Shapiro 2005) with permission.
Men and mice have two kinds of L chains, labeled Kappa (Lk) and Lambda (Ll). For each class of L chain, the germ line contains a single C region exon separated by introns from a set of four short join (J) region exons (JL1, JL2, JL3, JL4). Immediately upstream of the J region exons, no functional transcription signals are present. A considerable distance upstream are a set of about 100-300 V region exons preceded by suitable transcription signals (Vl1, Vl2…V li, V lj…V ly, V lz). A functional L chain coding sequence is constructed by joining one of the V regions to one of the J regions. The joined VJ exon can then be transcribed, and RNA splicing constructs a functional VJC coding sequence that can be translated into an L chain. It should be clear that different Vli –Jln and Vki –Jkn combinations can generate about 400-1200 different L chains of each class.We do not
fully understand the somatic hypermutation process or how it is
targeted, but a few informative details have been established (Papavasiliou
and Schatz 2002).
(i) The activated B cells synthesize an “activation-induced cytidine deaminase” (AID) enzyme that is essential for somatic hypermutation (Muramatsu, Kinoshita et al. 2000; Longerich, Basu et al. 2006). AID removes amino groups from C nucleotides in single-stranded DNA and converts them into Us, which are targets for base-excision repair. We hypothesize that AID action recruits repair functions, possibly including mutator polymerases (Cascalho, Wong et al. 1998; Diaz and Storb 2003; Seki, Gearhart et al. 2005; Goodman, Scharff et al. 2007; Liu and Schatz 2009), to participate in somatic hypermutation.
(ii) Somatic hypermutation depends on transcription from the immunoglobulin start signals and extends about 2 kb from the start site (Inlay, Gao et al. 2006). If other DNA sequences are placed downstream of the start site, then they are subject to hypermutation, and if the start site is moved next to the C region, then it is mutagenized. Thus, transcription is necessary for somatic hypermutation and acts in coordination with AID, almost certainly because the two DNA strands are separated during transcription (Ramiro, Stavropoulos et al. 2003).
(iii) A 9.8 kb sequence has recently been characterized that is essential for somatic hypermutation (Blagodatski, Batrak et al. 2009). This “diversification activator” (DIVAC) DNA segment lies downstream of the mutagenized V region and activates hypermutation when placed downstream of non-immunoglobulin loci in activated B cells. DIVAC appears to have a complex structure, and its mechanism of stimulating hypermutation is unknown. Since we know that AID works broadly around the genome in activated B cells, we may speculate that DIVAC somehow blocks the error-free repair that prevents hypermutation outside the immunoglobulin loci (Liu and Schatz 2009). The existence of DIVAC provides at least the beginnings of an explanation for the restriction of somatic hypermutation to immunoglobulin V region sequences.
The theoretical significance of somatic hypermutation is two fold, First, it demonstrates that cells can coordinate transcription signals, DNA-modifying activities like AID, and signals such as the DIVAC element to target highly localized and functionally relevant mutagenesis to a specific genome locus. This capacity has long been denied in conventional evolutionary thinking, and most evolutionists would treat somatic hypermutation as extraordinary and not of general significance. Nonetheless, the fact remains that living cells can, and do, carry out targeted mutagenesis. Consequently, this capability has to be incorporated into evolutionary thinking. The second significance of somatic hypermutation is that it demonstrates that rapid protein evolution occurs in two steps: (i) initial invention of the antigen-binding molecule by V(D)J joining and (ii) subsequent fine-tuning by targeted mutagenesis. This switch between macroevolution to produce something totally novel followed by microevolution to make the novelty work more efficiently may well be paradigmatic for evolution in general.
The conceptual importance of class switching lies in the demonstration that DS breaks can be targeted by lymphokine-directed transcription (Honjo, Kinoshita et al. 2002; Sellars, Reina-San-Martin et al.). The integration of intercellular signaling and the transcription it controls with DS breaks and NHEJ provides us with a detailed molecular mechanism for the way communication molecules can direct DNA rearrangements to particular regions of the genome. This is yet another capacity for controlling natural genetic engineering functions that has to be incorporated into our ideas about the genome restructuring tools available for evolution.
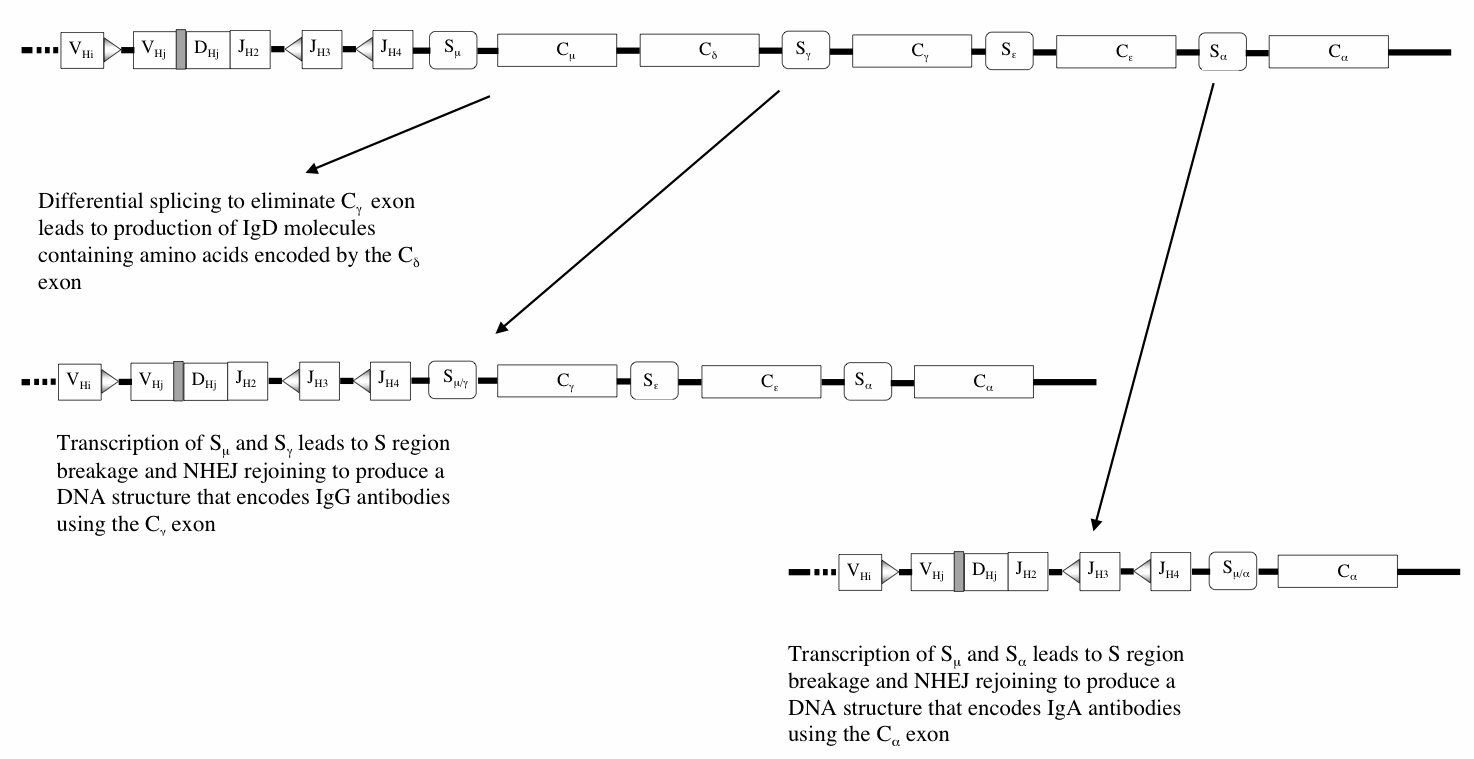
Class switch recombination, from (Shapiro 2005) with permission.
REFERENCES
Agrawal, A., Q. M. Eastman, et al. (1998). "Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system." Nature 394(6695): 744-751. http://www.ncbi.nlm.nih.gov/pubmed/9723614.
Aguilar, P. S., M. G. Heiman, et al. (2010). "Structure of sterol aliphatic chains affects yeast cell shape and cell fusion during mating." Proc Natl Acad Sci U S A 107(9): 4170-4175. http://www.ncbi.nlm.nih.gov/pubmed/20150508.
Alon, U. (2007). "Network motifs: theory and experimental approaches." Nat Rev Genet 8(6): 450-461. http://www.ncbi.nlm.nih.gov/pubmed/17510665.
Amoutzias, G. D., D. L. Robertson, et al. (2004). "Convergent evolution of gene networks by single-gene duplications in higher eukaryotes." EMBO Rep 5(3): 274-279. http://www.ncbi.nlm.nih.gov/pubmed/14968135.
Aravind, L., V. Anantharaman, et al. (2009). "Apprehending multicellularity: regulatory networks, genomics, and evolution." Birth Defects Res C Embryo Today 87(2): 143-164. http://www.ncbi.nlm.nih.gov/pubmed/19530132.
Arditti, R. R., J. G. Scaife, et al. (1968). "The nature of mutants in the lac promoter region." J Mol Biol 38(3): 421-426. http://www.ncbi.nlm.nih.gov/pubmed/4887878.
Arkowitz, R. A. (2009). "Chemical gradients and chemotropism in yeast." Cold Spring Harb Perspect Biol 1(2): a001958. http://www.ncbi.nlm.nih.gov/pubmed/20066086.
Arsene, F., T. Tomoyasu, et al. (2000). "The heat shock response of Escherichia coli." Int J Food Microbiol 55(1-3): 3-9. http://www.ncbi.nlm.nih.gov/pubmed/10791710.
Balaji, S. and L. Aravind (2007). "The two faces of short-range evolutionary dynamics of regulatory modes in bacterial transcriptional regulatory networks." Bioessays 29(7): 625-629. http://www.ncbi.nlm.nih.gov/pubmed/17563073.
Ben-Tabou de-Leon, S. and E. H. Davidson (2007). "Gene regulation: gene control network in development." Annu Rev Biophys Biomol Struct 36: 191. http://www.ncbi.nlm.nih.gov/pubmed/17291181.
Ben-Tabou de-Leon, S. and E. H. Davidson (2009). "Modeling the dynamics of transcriptional gene regulatory networks for animal development." Dev Biol 325(2): 317-328. http://www.ncbi.nlm.nih.gov/pubmed/19028486.
Benedict, C. L., S. Gilfillan, et al. (2000). "Terminal deoxynucleotidyl transferase and repertoire development." Immunol Rev 175: 150-157. http://www.ncbi.nlm.nih.gov/pubmed/10933600.
Bentolila, L. A., G. E. Wu, et al. (1997). "Constitutive expression of terminal deoxynucleotidyl transferase in transgenic mice is sufficient for N region diversity to occur at any Ig locus throughout B cell differentiation." J Immunol 158(2): 715-723. http://www.ncbi.nlm.nih.gov/pubmed/8992987.
Blagodatski, A., V. Batrak, et al. (2009). "A cis-acting diversification activator both necessary and sufficient for AID-mediated hypermutation." PLoS Genet 5(1): e1000332. http://www.ncbi.nlm.nih.gov/pubmed/19132090.
Blumer, K. J., J. E. Reneke, et al. (1988). "The STE2 gene product is the ligand-binding component of the alpha-factor receptor of Saccharomyces cerevisiae." J Biol Chem 263(22): 10836-10842. http://www.ncbi.nlm.nih.gov/pubmed/2839507.
Boboila, C., C. Yan, et al. (2010). "Alternative end-joining catalyzes class switch recombination in the absence of both Ku70 and DNA ligase 4." J Exp Med 207(2): 417-427. http://www.ncbi.nlm.nih.gov/pubmed/20142431.
Bonn, S. and E. E. Furlong (2008). "cis-Regulatory networks during development: a view of Drosophila." Curr Opin Genet Dev 18(6): 513-520. http://www.ncbi.nlm.nih.gov/pubmed/18929653.
Borok, M. J., D. A. Tran, et al. (2010). "Dissecting the regulatory switches of development: lessons from enhancer evolution in Drosophila." Development 137(1): 5-13. http://www.ncbi.nlm.nih.gov/pubmed/20023155.
Brandt, V. L. and D. B. Roth (2004). "V(D)J recombination: how to tame a transposase." Immunol Rev 200: 249-260. http://www.ncbi.nlm.nih.gov/pubmed/15242410.
Butler, J. E., Y. Zhao, et al. (2009). "Immunoglobulins, antibody repertoire and B cell development." Dev Comp Immunol 33(3): 321-333. http://www.ncbi.nlm.nih.gov/pubmed/18804488.
Calissano, C., D. Modiano, et al. (2003). "IgE antibodies to Plasmodium falciparum and severity of malaria in children of one ethnic group living in Burkina Faso." Am J Trop Med Hyg 69(1): 31-35. http://www.ncbi.nlm.nih.gov/pubmed/12932093.
Cascalho, M., J. Wong, et al. (1998). "Mismatch repair co-opted by hypermutation." Science 279(5354): 1207-1210. http://www.ncbi.nlm.nih.gov/pubmed/9469811.
Chatterji, M., C. L. Tsai, et al. (2004). "New concepts in the regulation of an ancient reaction: transposition by RAG1/RAG2." Immunol Rev 200: 261-271. http://www.ncbi.nlm.nih.gov/pubmed/15242411.
Chen, R. E. and J. Thorner (2007). "Function and regulation in MAPK signaling pathways: lessons learned from the yeast Saccharomyces cerevisiae." Biochim Biophys Acta 1773(8): 1311-1340. http://www.ncbi.nlm.nih.gov/pubmed/17604854.
Chenevert, J. (1994). "Cell polarization directed by extracellular cues in yeast." Mol Biol Cell 5(11): 1169-1175. http://www.ncbi.nlm.nih.gov/pubmed/7865882.
Chenevert, J., N. Valtz, et al. (1994). "Identification of genes required for normal pheromone-induced cell polarization in Saccharomyces cerevisiae." Genetics 136(4): 1287-1296. http://www.ncbi.nlm.nih.gov/pubmed/8013906.
Chou, S., S. Lane, et al. (2006). "Regulation of mating and filamentation genes by two distinct Ste12 complexes in Saccharomyces cerevisiae." Mol Cell Biol 26(13): 4794-4805. http://www.ncbi.nlm.nih.gov/pubmed/16782869.
Clatworthy, A. E., M. A. Valencia, et al. (2003). "V(D)J recombination and RAG-mediated transposition in yeast." Mol Cell 12(2): 489-499. http://www.ncbi.nlm.nih.gov/pubmed/14536087.
Cooper, M. B., M. Loose, et al. (2009). "The evolutionary influence of binding site organisation on gene regulatory networks." Biosystems 96(2): 185-193. http://www.ncbi.nlm.nih.gov/pubmed/19428984.
Corneo, B., R. L. Wendland, et al. (2007). "Rag mutations reveal robust alternative end joining." Nature 449(7161): 483-486. http://www.ncbi.nlm.nih.gov/pubmed/17898768.
Crasnier-Mednansky, M. (2008). "Is there any role for cAMP-CRP in carbon catabolite repression of the Escherichia coli lac operon?" Nat Rev Microbiol 6(12): 954; author reply 954. http://www.ncbi.nlm.nih.gov/pubmed/18955986.
Daniel, J. A., M. A. Santos, et al. (2010). "PTIP Promotes Chromatin Changes Critical for Immunoglobulin Class Switch Recombination." Science 329(5994): 917-923. http://www.ncbi.nlm.nih.gov/pubmed/20671152.
Davidson, E. H. and D. H. Erwin (2006). "Gene regulatory networks and the evolution of animal body plans." Science 311(5762): 796-800. http://www.ncbi.nlm.nih.gov/pubmed/16469913.
De Crombrugghe, B., B. Chen, et al. (1971). "Lac DNA, RNA polymerase and cyclic AMP receptor protein, cyclic AMP, lac repressor and inducer are the essential elements for controlled lac transcription." Nat New Biol 231(22): 139-142. http://www.ncbi.nlm.nih.gov/pubmed/4326354.
De Crombrugghe, B., R. L. Perlman, et al. (1969). "Regulation of inducible enzyme synthesis in Escherichia coli by cyclic adenosine 3', 5'-monophosphate." J Biol Chem 244(21): 5828-5835. http://www.ncbi.nlm.nih.gov/pubmed/4310825.
De Crombrugghe, E., B. Chen, et al. (1971). "Role of cyclic adenosine 3',5'-monophosphate and the cyclic adenosine 3',5'-monophosphate receptor protein in the initiation of lac transcription." J Biol Chem 246(23): 7343-7348. http://www.ncbi.nlm.nih.gov/pubmed/4333322.
De Robertis, E. M., G. Oliver, et al. (1990). "Homeobox genes and the vertebrate body plan." Sci Am 263(1): 46-52. http://www.ncbi.nlm.nih.gov/pubmed/1973846.
Diaz, M. and U. Storb (2003). "A novel cytidine deaminase AIDs in the delivery of error-prone polymerases to immunoglobulin genes." DNA Repair (Amst) 2(5): 623-627. http://www.ncbi.nlm.nih.gov/pubmed/12713818.
Dinkelmann, M., E. Spehalski, et al. (2009). "Multiple functions of MRN in end-joining pathways during isotype class switching." Nat Struct Mol Biol 16(8): 808-813. http://www.ncbi.nlm.nih.gov/pubmed/19633670.
Dohlman, H. G. (2002). "G proteins and pheromone signaling." Annu Rev Physiol 64: 129-152. http://www.ncbi.nlm.nih.gov/pubmed/11826266.
Dohlman, H. G. and J. E. Slessareva (2006). "Pheromone signaling pathways in yeast." Sci STKE 2006(364): cm6. http://www.ncbi.nlm.nih.gov/pubmed/17148787.
Dolan, J. W. (1996). "Novel aspects of pheromone-induced cell-cycle arrest in yeast." Curr Genet 30(6): 469-475. http://www.ncbi.nlm.nih.gov/pubmed/8939807.
Duarte, J., P. Deshpande, et al. (2007). "Total and functional parasite specific IgE responses in Plasmodium falciparum-infected patients exhibiting different clinical status." Malar J 6: 1. http://www.ncbi.nlm.nih.gov/pubmed/17204149.
Erb, K. J. (2007). "Helminths, allergic disorders and IgE-mediated immune responses: where do we stand?" Eur J Immunol 37(5): 1170-1173. http://www.ncbi.nlm.nih.gov/pubmed/17447233.
Erwin, D. H. and E. H. Davidson (2009). "The evolution of hierarchical gene regulatory networks." Nat Rev Genet 10(2): 141-148. http://www.ncbi.nlm.nih.gov/pubmed/19139764.
Esch, R. K., Y. Wang, et al. (2006). "Pheromone-induced degradation of Ste12 contributes to signal attenuation and the specificity of developmental fate." Eukaryot Cell 5(12): 2147-2160. http://www.ncbi.nlm.nih.gov/pubmed/17041188.
Fagarasan, S. (2008). "Evolution, development, mechanism and function of IgA in the gut." Curr Opin Immunol 20(2): 170-177. http://www.ncbi.nlm.nih.gov/pubmed/18456485.
Fagarasan, S. and T. Honjo (2003). "Intestinal IgA synthesis: regulation of front-line body defences." Nat Rev Immunol 3(1): 63-72. http://www.ncbi.nlm.nih.gov/pubmed/12511876.
Fitzsimmons, C. M., R. McBeath, et al. (2007). "Factors affecting human IgE and IgG responses to allergen-like Schistosoma mansoni antigens: Molecular structure and patterns of in vivo exposure." Int Arch Allergy Immunol 142(1): 40-50. http://www.ncbi.nlm.nih.gov/pubmed/17019080.
Friedman, B. E., J. S. Olson, et al. (1977). "Interaction of lac repressor with inducer, kinetic and equilibrium measurements." J Mol Biol 111(1): 27-39. http://www.ncbi.nlm.nih.gov/pubmed/16140.
Fugmann, S. D. (2001). "RAG1 and RAG2 in V(D)J recombination and transposition." Immunol Res 23(1): 23-39. http://www.ncbi.nlm.nih.gov/pubmed/11417858.
Gellert, M. (1997). "Recent advances in understanding V(D)J recombination." Adv Immunol 64: 39-64. http://www.ncbi.nlm.nih.gov/pubmed/9100979.
Gonzalez Perez, A. D., E. Gonzalez Gonzalez, et al. (2008). "Impact of Transcription Units rearrangement on the evolution of the regulatory network of gamma-proteobacteria." BMC Genomics 9: 128. http://www.ncbi.nlm.nih.gov/pubmed/18366643.
Goodman, M. F., M. D. Scharff, et al. (2007). "AID-initiated purposeful mutations in immunoglobulin genes." Adv Immunol 94: 127-155. http://www.ncbi.nlm.nih.gov/pubmed/17560274.
Gramzow, L., M. S. Ritz, et al. (2010). "On the origin of MADS-domain transcription factors." Trends Genet 26(4): 149-153. http://www.ncbi.nlm.nih.gov/pubmed/20219261.
Grimm, S. L. and S. K. Nordeen (1999). "A composite enhancer element directing tissue-specific expression of mouse mammary tumor virus requires both ubiquitous and tissue-restricted factors." J Biol Chem 274(18): 12790-12796. http://www.ncbi.nlm.nih.gov/pubmed/10212264.
Gustin, M. C., J. Albertyn, et al. (1998). "MAP kinase pathways in the yeast Saccharomyces cerevisiae." Microbiol Mol Biol Rev 62(4): 1264-1300. http://www.ncbi.nlm.nih.gov/pubmed/9841672.
Hackney, J. A., S. Misaghi, et al. (2009). "DNA targets of AID evolutionary link between antibody somatic hypermutation and class switch recombination." Adv Immunol 101: 163-189. http://www.ncbi.nlm.nih.gov/pubmed/19231595.
Haerty, W., C. Artieri, et al. (2008). "Comparative analysis of function and interaction of transcription factors in nematodes: extensive conservation of orthology coupled to rapid sequence evolution." BMC Genomics 9: 399. http://www.ncbi.nlm.nih.gov/pubmed/18752680.
Harwood, J. P., C. Gazdar, et al. (1976). "Involvement of the glucose enzymes II of the sugar phosphotransferase system in the regulation of adenylate cyclase by glucose in Escherichia coli." J Biol Chem 251(8): 2462-2468. http://www.ncbi.nlm.nih.gov/pubmed/177417.
Hengge-Aronis, R. (2002). "Recent insights into the general stress response regulatory network in Escherichia coli." J Mol Microbiol Biotechnol 4(3): 341-346. http://www.ncbi.nlm.nih.gov/pubmed/11931567.
Hickey, A. J., E. Conway de Macario, et al. (2002). "Transcription in the archaea: basal factors, regulation, and stress-gene expression." Crit Rev Biochem Mol Biol 37(6): 537-599. http://www.ncbi.nlm.nih.gov/pubmed/12540196.
Hinman, V. F., A. Nguyen, et al. (2007). "Caught in the evolutionary act: precise cis-regulatory basis of difference in the organization of gene networks of sea stars and sea urchins." Dev Biol 312(2): 584-595. http://www.ncbi.nlm.nih.gov/pubmed/17956756.
Hinman, V. F., K. A. Yankura, et al. (2009). "Evolution of gene regulatory network architectures: examples of subcircuit conservation and plasticity between classes of echinoderms." Biochim Biophys Acta 1789(4): 326-332. http://www.ncbi.nlm.nih.gov/pubmed/19284985.
Hiom, K., M. Melek, et al. (1998). "DNA transposition by the RAG1 and RAG2 proteins: a possible source of oncogenic translocations." Cell 94(4): 463-470. http://www.ncbi.nlm.nih.gov/pubmed/9727489.
Ho, M. C., H. Johnsen, et al. (2009). "Functional evolution of cis-regulatory modules at a homeotic gene in Drosophila." PLoS Genet 5(11): e1000709. http://www.ncbi.nlm.nih.gov/pubmed/19893611.
Honjo, T., K. Kinoshita, et al. (2002). "Molecular mechanism of class switch recombination: linkage with somatic hypermutation." Annu Rev Immunol 20: 165-196. http://www.ncbi.nlm.nih.gov/pubmed/11861601.
Hu, B., W. J. Rappel, et al. (2009). "Mechanisms and constraints on yeast MAPK signaling specificity." Biophys J 96(12): 4755-4763. http://www.ncbi.nlm.nih.gov/pubmed/19527636.
Huber, R. E., G. Kurz, et al. (1976). "A quantitation of the factors which affect the hydrolase and transgalactosylase activities of beta-galactosidase (E. coli) on lactose." Biochemistry 15(9): 1994-2001. http://www.ncbi.nlm.nih.gov/pubmed/5122.
Huber, R. E., K. Wallenfels, et al. (1975). "The action of beta-galactosidase (Escherichia coli) on allolactose." Can J Biochem 53(9): 1035-1038. http://www.ncbi.nlm.nih.gov/pubmed/241475.
Hughes, A. L. and R. Friedman (2007). "Sharing of transcription factors after gene duplication in the yeast Saccharomyces cerevisiae." Genetica 129(3): 301-308. http://www.ncbi.nlm.nih.gov/pubmed/16897462.
Inada, T., K. Kimata, et al. (1996). "Mechanism responsible for glucose-lactose diauxie in Escherichia coli: challenge to the cAMP model." Genes Cells 1(3): 293-301. http://www.ncbi.nlm.nih.gov/pubmed/9133663.
Inlay, M. A., H. H. Gao, et al. (2006). "Roles of the Ig kappa light chain intronic and 3' enhancers in Igk somatic hypermutation." J Immunol 177(2): 1146-1151. http://www.ncbi.nlm.nih.gov/pubmed/16818772.
Jacob, F. and J. Monod (1961). "Genetic regulatory mechanisms in the synthesis of proteins." J Mol Biol 3: 318-356. http://www.ncbi.nlm.nih.gov/pubmed/13718526\.
Jaeger, S. A., E. T. Chan, et al. (2010). "Conservation and regulatory associations of a wide affinity range of mouse transcription factor binding sites." Genomics 95(4): 185-195. http://www.ncbi.nlm.nih.gov/pubmed/20079828.
Jolly, C. J., A. J. Cook, et al. (2008). "Fixing DNA breaks during class switch recombination." J Exp Med 205(3): 509-513. http://www.ncbi.nlm.nih.gov/pubmed/18332183\.
Jones, J. M. and M. Gellert (2004). "The taming of a transposon: V(D)J recombination and the immune system." Immunol Rev 200: 233-248. http://www.ncbi.nlm.nih.gov/pubmed/15242409.
Josefsson, L. G. (1999). "Evidence for kinship between diverse G-protein coupled receptors." Gene 239(2): 333-340. http://www.ncbi.nlm.nih.gov/pubmed/10548735.
Jung, D., C. Giallourakis, et al. (2006). "Mechanism and control of V(D)J recombination at the immunoglobulin heavy chain locus." Annu Rev Immunol 24: 541-570. http://www.ncbi.nlm.nih.gov/pubmed/16551259.
Kimata, K., H. Takahashi, et al. (1997). "cAMP receptor protein-cAMP plays a crucial role in glucose-lactose diauxie by activating the major glucose transporter gene in Escherichia coli." Proc Natl Acad Sci U S A 94(24): 12914-12919. http://www.ncbi.nlm.nih.gov/pubmed/9371775.
Kinoshita, K. and T. Honjo (2001). "Linking class-switch recombination with somatic hypermutation." Nat Rev Mol Cell Biol 2(7): 493-503. http://www.ncbi.nlm.nih.gov/pubmed/11433363.
Kirkman-Correia, C., I. L. Stroke, et al. (1993). "Functional domains of the yeast STE12 protein, a pheromone-responsive transcriptional activator." Mol Cell Biol 13(6): 3765-3772. http://www.ncbi.nlm.nih.gov/pubmed/8497278.
Klauck, E., A. Typas, et al. (2007). "The sigmaS subunit of RNA polymerase as a signal integrator and network master regulator in the general stress response in Escherichia coli." Sci Prog 90(Pt 2-3): 103-127. http://www.ncbi.nlm.nih.gov/pubmed/17725229.
Koelle, M. R. (2006). "Heterotrimeric G protein signaling: Getting inside the cell." Cell 126(1): 25-27. http://www.ncbi.nlm.nih.gov/pubmed/16839871.
Kosko, B. and S. Isaka (1993). "Fuzzy Logic." Scientific American 269(1): 76-81. .
Kuhlman, T., Z. Zhang, et al. (2007). "Combinatorial transcriptional control of the lactose operon of Escherichia coli." Proc Natl Acad Sci U S A 104(14): 6043-6048. http://www.ncbi.nlm.nih.gov/pubmed/17376875.
Kundig, W., S. Ghosh, et al. (1964). "Phosphate Bound to Histidine in a Protein as an Intermediate in a Novel Phospho-Transferase System." Proc Natl Acad Sci U S A 52: 1067-1074. http://www.ncbi.nlm.nih.gov/pubmed/14224387.
Kusari, A. B., D. M. Molina, et al. (2004). "A conserved protein interaction network involving the yeast MAP kinases Fus3 and Kss1." J Cell Biol 164(2): 267-277. http://www.ncbi.nlm.nih.gov/pubmed/14734536.
Lee, B. N. and E. A. Elion (1999). "The MAPKKK Ste11 regulates vegetative growth through a kinase cascade of shared signaling components." Proc Natl Acad Sci U S A 96(22): 12679-12684. http://www.ncbi.nlm.nih.gov/pubmed/10535982.
Lee, G. S., M. B. Neiditch, et al. (2004). "RAG proteins shepherd double-strand breaks to a specific pathway, suppressing error-prone repair, but RAG nicking initiates homologous recombination." Cell 117(2): 171-184. http://www.ncbi.nlm.nih.gov/pubmed/15084256.
Leoratti, F. M., R. R. Durlacher, et al. (2008). "Pattern of humoral immune response to Plasmodium falciparum blood stages in individuals presenting different clinical expressions of malaria." Malar J 7: 186. http://www.ncbi.nlm.nih.gov/pubmed/18816374.
Lesage, G. and H. Bussey (2006). "Cell wall assembly in Saccharomyces cerevisiae." Microbiol Mol Biol Rev 70(2): 317-343. http://www.ncbi.nlm.nih.gov/pubmed/16760306.
Levine, M. and E. H. Davidson (2005). "Gene regulatory networks for development." Proc Natl Acad Sci U S A 102(14): 4936-4942. http://www.ncbi.nlm.nih.gov/pubmed/15788537.
Li, E. and E. H. Davidson (2009). "Building developmental gene regulatory networks." Birth Defects Res C Embryo Today 87(2): 123-130. http://www.ncbi.nlm.nih.gov/pubmed/19530131.
Liu, M. and D. G. Schatz (2009). "Balancing AID and DNA repair during somatic hypermutation." Trends Immunol\ 30(4): 173-181. http://www.ncbi.nlm.nih.gov/pubmed/19303358.
Longerich, S., U. Basu, et al. (2006). "AID in somatic hypermutation and class switch recombination." Curr Opin Immunol 18(2): 164-174. http://www.ncbi.nlm.nih.gov/pubmed/16464563.
MacArthur, S., X. Y. Li, et al. (2009). "Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions." Genome Biol 10(7): R80. http://www.ncbi.nlm.nih.gov/pubmed/19627575.
Madan Babu, M., S. Balaji, et al. (2007). "General trends in the evolution of prokaryotic transcriptional regulatory networks." Genome Dyn 3: 66-80. http://www.ncbi.nlm.nih.gov/pubmed/18753785.
Madan Babu, M., S. A. Teichmann, et al. (2006). "Evolutionary dynamics of prokaryotic transcriptional regulatory networks." J Mol Biol 358(2): 614-633. http://www.ncbi.nlm.nih.gov/pubmed/16530225.
Mangan, S. and U. Alon (2003). "Structure and function of the feed-forward loop network motif." Proc Natl Acad Sci U S A 100(21): 11980-11985. http://www.ncbi.nlm.nih.gov/pubmed/14530388.
Matthews, A. G. and M. A. Oettinger (2009). "Regulation of RAG transposition." Adv Exp Med Biol 650: 16-31. http://www.ncbi.nlm.nih.gov/pubmed/19731798.
McCullough, K. C. and A. Summerfield (2005). "Basic concepts of immune response and defense development." Ilar J 46(3): 230-240. http://www.ncbi.nlm.nih.gov/pubmed/15953831.
Mishra, N. S., R. Tuteja, et al. (2006). "Signaling through MAP kinase networks in plants." Arch Biochem Biophys 452(1): 55-68. http://www.ncbi.nlm.nih.gov/pubmed/16806044.
Monod, J. and M. Cohn (1952). "[Biosynthesis induced by enzymes; enzymatic adaptation.]." Adv Enzymol Relat Subj Biochem 13: 67-119. http://www.ncbi.nlm.nih.gov/pubmed/14943665.
Muller, J., S. Oehler, et al. (1996). "Repression of lac promoter as a function of distance, phase and quality of an auxiliary lac operator." J Mol Biol 257(1): 21-29. http://www.ncbi.nlm.nih.gov/pubmed/8632456.
Muramatsu, M., K. Kinoshita, et al. (2000). "Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme." Cell 102(5): 553-563. http://www.ncbi.nlm.nih.gov/pubmed/11007474.
Narang, A. (2009). "cAMP does not have an important role in carbon catabolite repression of the Escherichia coli lac operon." Nat Rev Microbiol 7(3): 250. http://www.ncbi.nlm.nih.gov/pubmed/19230236.
Narang, A. (2009). "Quantitative effect and regulatory function of cyclic adenosine 5'-phosphate in Escherichia coli." J Biosci 34(3): 445-463. http://www.ncbi.nlm.nih.gov/pubmed/19805906.
Neiditch, M. B., G. S. Lee, et al. (2002). "The V(D)J recombinase efficiently cleaves and transposes signal joints." Mol Cell 9(4): 871-878. http://www.ncbi.nlm.nih.gov/pubmed/11983177.
Nick McElhinny, S. A. and D. A. Ramsden (2004). "Sibling rivalry: competition between Pol X family members in V(D)J recombination and general double strand break repair." Immunol Rev 200: 156-164. http://www.ncbi.nlm.nih.gov/pubmed/15242403.
Nisseley, S. P., W. B. Anderson, et al. (1971). "In vitro transcription of the gal operon requires cyclic adenosine monophosphate and cyclic adenosine monophosphate receptor protein." J Biol Chem 246(15): 4671-4678. http://www.ncbi.nlm.nih.gov/pubmed/4327324.
Nomoto, S., N. Nakayama, et al. (1990). "Regulation of the yeast pheromone response pathway by G protein subunits." Embo J 9(3): 691-696. http://www.ncbi.nlm.nih.gov/pubmed/2107073.
Nusslein-Volhard, C. (1996). "Gradients that organize embryo development." Sci Am 275(2): 54-55; 58-61. http://www.ncbi.nlm.nih.gov/pubmed/8693324.
Nusslein-Volhard, C. and E. Wieschaus (1980). "Mutations affecting segment number and polarity in Drosophila." Nature 287(5785): 795-801. http://www.ncbi.nlm.nih.gov/pubmed/6776413.
Paliwal, S., P. A. Iglesias, et al. (2007). "MAPK-mediated bimodal gene expression and adaptive gradient sensing in yeast." Nature 446(7131): 46-51. http://www.ncbi.nlm.nih.gov/pubmed/17310144.
Papavasiliou, F. N. and D. G. Schatz (2002). "Somatic hypermutation of immunoglobulin genes: merging mechanisms for genetic diversity." Cell 109 Suppl: S35-44. http://www.ncbi.nlm.nih.gov/pubmed/11983151.
Pastan, I. and R. Perlman (1970). "Cyclic adenosine monophosphate in bacteria." Science 169(943): 339-344. http://www.ncbi.nlm.nih.gov/pubmed/4317896.
Pastan, I. and R. L. Perlman (1972). "Regulation of gene transcription in Escherichia coli by cyclic AMP." Adv Cyclic Nucleotide Res 1: 11-16. http://www.ncbi.nlm.nih.gov/pubmed/4353161.
Perlman, R. L., B. De Crombrugghe, et al. (1969). "Cyclic AMP regulates catabolite and transient repression in E. coli." Nature 223(5208): 810-812. http://www.ncbi.nlm.nih.gov/pubmed/4307969.
Perlman, R. L. and I. Pastan (1968). "Regulation of beta-galactosidase synthesis in Escherichia coli by cyclic adenosine 3',5'-monophosphate." J Biol Chem 243(20): 5420-5427. http://www.ncbi.nlm.nih.gov/pubmed/4302785.
Peterkofsky, A. (1988). "Redistribution of phosphate pools and the regulation of Escherichia coli adenylate cyclase activity." Arch Biochem Biophys 265(2): 227-233. http://www.ncbi.nlm.nih.gov/pubmed/2844115.
Pfister, K., K. Turner, et al. (1983). "IgE production in rat fascioliasis." Parasite Immunol 5(6): 587-593. http://www.ncbi.nlm.nih.gov/pubmed/6657297.
Pinney, J. W., G. D. Amoutzias, et al. (2007). "Reconstruction of ancestral protein interaction networks for the bZIP transcription factors." Proc Natl Acad Sci U S A 104(51): 20449-20453. http://www.ncbi.nlm.nih.gov/pubmed/18077348.
Pirkkala, L., P. Nykanen, et al. (2001). "Roles of the heat shock transcription factors in regulation of the heat shock response and beyond." Faseb J 15(7): 1118-1131. http://www.ncbi.nlm.nih.gov/pubmed/11344080.
Ramiro, A. R., P. Stavropoulos, et al. (2003). "Transcription enhances AID-mediated cytidine deamination by exposing single-stranded DNA on the nontemplate strand." Nat Immunol 4(5): 452-456. http://www.ncbi.nlm.nih.gov/pubmed/12692548.
Roseman, S. (1969). "The transport of carbohydrates by a bacterial phosphotransferase system." J Gen Physiol 54(1): 138-184. http://www.ncbi.nlm.nih.gov/pubmed/19873641.
Saalmuller, A. (2006). "New understanding of immunological mechanisms." Vet Microbiol 117(1): 32-38. http://www.ncbi.nlm.nih.gov/pubmed/16701965.
Samaj, J., F. Baluska, et al. (2004). "From signal to cell polarity: mitogen-activated protein kinases as sensors and effectors of cytoskeleton dynamicity." J Exp Bot 55(395): 189-198. http://www.ncbi.nlm.nih.gov/pubmed/14673033.
Savery, N., V. Rhodius, et al. (1996). "Protein-protein interactions during transcription activation: the case of the Escherichia coli cyclic AMP receptor protein." Philos Trans R Soc Lond B Biol Sci 351(1339): 543-550. http://www.ncbi.nlm.nih.gov/pubmed/8735277.
Scaife, J. and J. R. Beckwith (1966). "Mutational alteration of the maximal level of Lac operon expression." Cold Spring Harb Symp Quant Biol 31: 403-408. http://www.ncbi.nlm.nih.gov/pubmed/4866390.
Schwacke, J. H. and E. O. Voit (2007). "The potential for signal integration and processing in interacting MAP kinase cascades." J Theor Biol 246(4): 604-620. http://www.ncbi.nlm.nih.gov/pubmed/17337011.
Seki, M., P. J. Gearhart, et al. (2005). "DNA polymerases and somatic hypermutation of immunoglobulin genes." EMBO Rep 6(12): 1143-1148. http://www.ncbi.nlm.nih.gov/pubmed/16319960.
Sellars, M., B. Reina-San-Martin, et al. (2009). "Ikaros controls isotype selection during immunoglobulin class switch recombination." J Exp Med 206(5): 1073-1087. http://www.ncbi.nlm.nih.gov/pubmed/19414557.
Shan, H., L. Zahn, et al. (2009). "Evolution of plant MADS box transcription factors: evidence for shifts in selection associated with early angiosperm diversification and concerted gene duplications." Mol Biol Evol 26(10): 2229-2244. http://www.ncbi.nlm.nih.gov/pubmed/19578156.
Shapiro, J. A. (2005). "A 21st century view of evolution: genome system architecture, repetitive DNA, and natural genetic engineering." Gene 345(1): 91-100. http://www.ncbi.nlm.nih.gov/pubmed/15716117.
Silverstone, A. E., B. Magasanik, et al. (1969). "Catabolite sensitive site of the lac operon." Nature 221(5185): 1012-1014. http://www.ncbi.nlm.nih.gov/pubmed/4886742.
Simoni, R. D., M. Levinthal, et al. (1967). "Genetic evidence for the role of a bacterial phosphotransferase system in sugar transport." Proc Natl Acad Sci U S A 58(5): 1963-1970. http://www.ncbi.nlm.nih.gov/pubmed/4866983.
Slessareva, J. E. and H. G. Dohlman (2006). "G protein signaling in yeast: new components, new connections, new compartments." Science 314(5804): 1412-1413. http://www.ncbi.nlm.nih.gov/pubmed/17138892.
Song, D., J. W. Dolan, et al. (1991). "Pheromone-dependent phosphorylation of the yeast STE12 protein correlates with transcriptional activation." Genes Dev 5(5): 741-750. http://www.ncbi.nlm.nih.gov/pubmed/2026326.
Sprague, G. F., Jr. (1991). "Signal transduction in yeast mating: receptors, transcription factors, and the kinase connection." Trends Genet 7(11-12): 393-398. http://www.ncbi.nlm.nih.gov/pubmed/1668192.
Stock, J. B., E. B. Waygood, et al. (1982). "Sugar transport by the bacterial phosphotransferase system. The glucose receptors of the Salmonella typhimurium phosphotransferase system." J Biol Chem 257(23): 14543-14552. http://www.ncbi.nlm.nih.gov/pubmed/6292227.
Tangteerawatana, P., S. M. Montgomery, et al. (2007). "Differential regulation of IgG subclasses and IgE antimalarial antibody responses in complicated and uncomplicated Plasmodium falciparum malaria." Parasite Immunol 29(9): 475-483. http://www.ncbi.nlm.nih.gov/pubmed/17727571.
Tjian, R. (1995). "Molecular Machines that Control Genes." Scientific American 272(2): 54-61. http://www.ncbi.nlm.nih.gov/pubmed/ 7817187.
Tuaillon, N. and J. D. Capra (1998). "Use of D gene segments with irregular spacers in terminal deoxynucleotidyltransferase (TdT)+/+ and TdT-/- mice carrying a human Ig heavy chain transgenic minilocus." Proc Natl Acad Sci U S A 95(4): 1703-1708. http://www.ncbi.nlm.nih.gov/pubmed/9465080.
Tuaillon, N. and J. D. Capra (2000). "Evidence that terminal deoxynucleotidyltransferase expression plays a role in Ig heavy chain gene segment utilization." J Immunol 164(12): 6387-6397. http://www.ncbi.nlm.nih.gov/pubmed/10843694.
Turner, C. L. and R. E. Huber (1977). "Differential binding of allolactose anomers to the lactose repressor of Escherichia coli." J Mol Biol 115(2): 195-199. http://www.ncbi.nlm.nih.gov/pubmed/338908.
van Drogen, F. and M. Peter (2001). "MAP kinase dynamics in yeast." Biol Cell 93(1-2): 63-70. http://www.ncbi.nlm.nih.gov/pubmed/11730324.
van Gent, D. C. and M. van der Burg (2007). "Non-homologous end-joining, a sticky affair." Oncogene 26(56): 7731-7740. http://www.ncbi.nlm.nih.gov/pubmed/18066085.
Veitia, R. A. (2008). "One thousand and one ways of making functionally similar transcriptional enhancers." Bioessays 30(11-12): 1052-1057. http://www.ncbi.nlm.nih.gov/pubmed/18937349.
Wallrath, L. L., Q. Lu, et al. (1994). "Architectural variations of inducible eukaryotic promoters: preset and remodeling chromatin structures." Bioessays 16(3): 165-170. http://www.ncbi.nlm.nih.gov/pubmed/8166669.
Wang, J. H., M. Gostissa, et al. (2009). "Mechanisms promoting translocations in editing and switching peripheral B cells." Nature 460(7252): 231-236. http://www.ncbi.nlm.nih.gov/pubmed/19587764.
Wang, Y. and H. G. Dohlman (2002). "Pheromone-dependent ubiquitination of the mitogen-activated protein kinase kinase Ste7." J Biol Chem 277(18): 15766-15772. http://www.ncbi.nlm.nih.gov/pubmed/11864977.
Wang, Y. and H. G. Dohlman (2006). "Pheromone-regulated sumoylation of transcription factors that mediate the invasive to mating developmental switch in yeast." J Biol Chem 281(4): 1964-1969. http://www.ncbi.nlm.nih.gov/pubmed/16306045.
Wang, Y. and H. G. Dohlman (2006). "Regulation of G protein and mitogen-activated protein kinase signaling by ubiquitination: insights from model organisms." Circ Res 99(12): 1305-1314. http://www.ncbi.nlm.nih.gov/pubmed/17158346.
Watanabe, N., F. Bruschi, et al. (2005). "IgE: a question of protective immunity in Trichinella spiralis infection." Trends Parasitol 21(4): 175-178. http://www.ncbi.nlm.nih.gov/pubmed/15780839.
Weterings, E. and D. J. Chen (2008). "The endless tale of non-homologous end-joining." Cell Res 18(1): 114-124. http://www.ncbi.nlm.nih.gov/pubmed/18166980.
Wu, R., C. P. Bahl, et al. (1978). "Lactose operator--repressor interaction." Curr Top Cell Regul 13: 137-178. http://www.ncbi.nlm.nih.gov/pubmed/352620.
Yan, C. T., C. Boboila, et al. (2007). "IgH class switching and translocations use a robust non-classical end-joining pathway." Nature 449(7161): 478-482. http://www.ncbi.nlm.nih.gov/pubmed/17713479.
Yin, D., E. Shumay, et al. (2006). "Yeast Ste2 receptors as tools for study of mammalian protein kinases and adaptors involved in receptor trafficking." J Mol Signal 1: 2. http://www.ncbi.nlm.nih.gov/pubmed/17224079.
Zan, H. and P. Casali (2008). "AID- and Ung-dependent generation of staggered double-strand DNA breaks in immunoglobulin class switch DNA recombination: a post-cleavage role for AID." Mol Immunol 46(1): 45-61. http://www.ncbi.nlm.nih.gov/pubmed/18760480.
Zan, H., A. Cerutti, et al. (1999). "Induction of Ig somatic hypermutation and class switching in a human monoclonal IgM+ IgD+ B cell line in vitro: definition of the requirements and modalities of hypermutation." J Immunol 162(6): 3437-3447. http://www.ncbi.nlm.nih.gov/pubmed/10092799.
Zhou, L., R. Mitra, et al. (2004). "Transposition of hAT elements links transposable elements and V(D)J recombination." Nature 432(7020): 995-1001. http://www.ncbi.nlm.nih.gov/pubmed/15616554.
Zinzen, R. P., C. Girardot, et al. (2009). "Combinatorial binding predicts spatio-temporal cis-regulatory activity." Nature 462(7269): 65-70. http://www.ncbi.nlm.nih.gov/pubmed/19890324.